Rows: 344
Columns: 8
$ species <fct> Adelie, Adelie, Adelie, Adelie, Adelie, Adelie, Adel…
$ island <fct> Torgersen, Torgersen, Torgersen, Torgersen, Torgerse…
$ bill_length_mm <dbl> 39.1, 39.5, 40.3, NA, 36.7, 39.3, 38.9, 39.2, 34.1, …
$ bill_depth_mm <dbl> 18.7, 17.4, 18.0, NA, 19.3, 20.6, 17.8, 19.6, 18.1, …
$ flipper_length_mm <int> 181, 186, 195, NA, 193, 190, 181, 195, 193, 190, 186…
$ body_mass_g <int> 3750, 3800, 3250, NA, 3450, 3650, 3625, 4675, 3475, …
$ sex <fct> male, female, female, NA, female, male, female, male…
$ year <int> 2007, 2007, 2007, 2007, 2007, 2007, 2007, 2007, 2007…Two-Way ANOVA Application
STAT 218 - Week 9, Lecture 2 Lab 9
March 12th, 2024
Two-Way ANOVA in R
Let’s Load Data Set
A Note for Assumption Checking
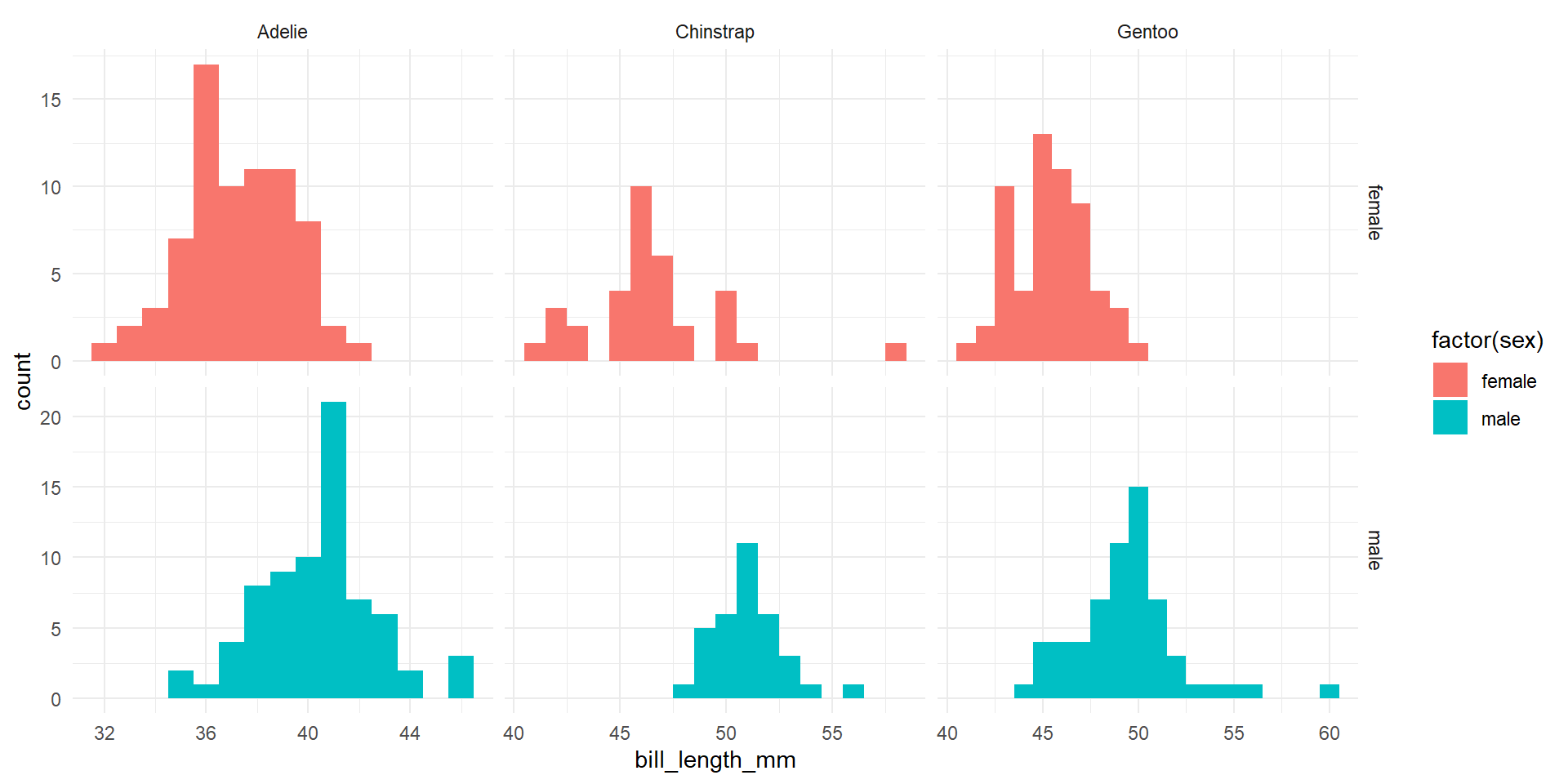
A Note for Assumption Checking
na.omit(penguins) |>
group_by(sex:species) |>
summarize(mean_bill_length = mean(bill_length_mm), sd_bill_length = sd(bill_length_mm))# A tibble: 6 × 3
`sex:species` mean_bill_length sd_bill_length
<fct> <dbl> <dbl>
1 female:Adelie 37.3 2.03
2 female:Chinstrap 46.6 3.11
3 female:Gentoo 45.6 2.05
4 male:Adelie 40.4 2.28
5 male:Chinstrap 51.1 1.56
6 male:Gentoo 49.5 2.72A Note for Assumption Checking
Two-Way ANOVA Function
Df Sum Sq Mean Sq F value Pr(>F)
sex 1 1175 1175 219.228 <2e-16 ***
species 2 6976 3488 650.479 <2e-16 ***
sex:species 2 24 12 2.284 0.103
Residuals 327 1753 5
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
11 observations deleted due to missingnessMain Effect 2
Interaction Effect
Tukey multiple comparisons of means
95% family-wise confidence level
Fit: aov(formula = bill_length_mm ~ sex + species + sex * species, data = penguins)
$`sex:species`
diff lwr upr p adj
male:Adelie-female:Adelie 3.132877 2.034137 4.2316165 0.0000000
female:Chinstrap-female:Adelie 9.315995 7.937732 10.6942588 0.0000000
male:Chinstrap-female:Adelie 13.836583 12.458320 15.2148470 0.0000000
female:Gentoo-female:Adelie 8.306259 7.138639 9.4738788 0.0000000
male:Gentoo-female:Adelie 12.216236 11.064727 13.3677453 0.0000000
female:Chinstrap-male:Adelie 6.183118 4.804855 7.5613821 0.0000000
male:Chinstrap-male:Adelie 10.703707 9.325443 12.0819703 0.0000000
female:Gentoo-male:Adelie 5.173382 4.005762 6.3410021 0.0000000
male:Gentoo-male:Adelie 9.083360 7.931851 10.2348686 0.0000000
male:Chinstrap-female:Chinstrap 4.520588 2.910622 6.1305547 0.0000000
female:Gentoo-female:Chinstrap -1.009736 -2.443514 0.4240412 0.3338130
male:Gentoo-female:Chinstrap 2.900241 1.479553 4.3209291 0.0000002
female:Gentoo-male:Chinstrap -5.530325 -6.964102 -4.0965471 0.0000000
male:Gentoo-male:Chinstrap -1.620347 -3.041035 -0.1996591 0.0149963
male:Gentoo-female:Gentoo 3.909977 2.692570 5.1273846 0.0000000An Important Note
- When interactions are present, the main effects of factors don’t have their usual interpretations.
- Because of this, we usually test for the presence of interactions first.
- If interactions are present, then we often stop the analysis at this stage and examine interaction effects.
- If no evidence for an interaction effect is found (i.e., if we do not reject \(H_0\)), then we proceed to testing the main effects of the individual factors.
- Because of this, we usually test for the presence of interactions first.
- So in this example, we ignored Main Effect 1 & Main Effect 2 and examine the Interaction Effect!